Nonlinear dimensionality reduction for manifold processing

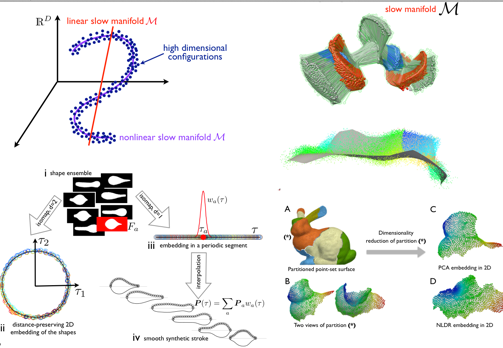
Simplifying a complex system in terms of a reduced set of coordinates is of great interest to gain insight and to accelerate computations. Most often, reduced models are a linear superposition of "modes" (e.g., PCA). Yet in some applications, the underlying geometry is nonlinear, leading to a very large number of linear modes compared to the intrinsic dimension. We are developing methods to turn statistical learning by nonlinear dimensionality reduction into a computational tool to perform calculations on the slow manifold of the system.
We have applied these methods to smooth point-set surface parametrization for thin shell analysis, to parametrize motility strokes of unicellular organisms, to model reduction in nonlinear elastodynamics, or to identify differentiable collective variables for enhanced sampling in molecular dynamics.
Publications:
Nonlinear manifold learning for model reduction in finite elastodynamics
Millán, D. and Arroyo, M.
Computer Methods in Applied Mechanics and Engineering, 261-262, 118-131 (2013)
http://dx.doi.org/10.1016/j.cma.2013.04.007
Reverse engineering the euglenoid movement
M. Arroyo, L. Heltai, D. Millán, A. DeSimone, Proceedings of the National Academy of Sciences USA, 109, 17874 (2012).
Nonlinear manifold learning for meshfree finite deformation thin shell analysis
D. Millán, A. Rosolen, M. Arroyo, International Journal for Numerical Methods in Engineering, DOI: 10.1002/nme.4403 (2012).
Monday, October 1, 2012