
COMPLIMB: A COMPUTATIONAL TOOL TO ELUCIDATE THE MECHANOBIOLOGICAL REGULATION OF LIMB DEVELOPMENT
10 July, 2019 @ 2:30 pm - 3:30 pm
 Ester Comellas is a postdoctoral researcher at the Shefelbine Lab in Northeastern University (Boston, US) since January 2019. Her main research interest is the computational modeling of growth and remodeling in soft tissues. Having focused on numerics and mechanics during her initial formation, she is now keen on learning more about the biological aspects of these phenomena. Recently awarded a Marie Curie fellowship, she will work closely with developmental biologists the following 2 years to elucidate the mechanobiology of joint morphogenesis, and then move back to the UPC in Barcelona to complete the CompLimb project in LaCàN.
Ester Comellas is a postdoctoral researcher at the Shefelbine Lab in Northeastern University (Boston, US) since January 2019. Her main research interest is the computational modeling of growth and remodeling in soft tissues. Having focused on numerics and mechanics during her initial formation, she is now keen on learning more about the biological aspects of these phenomena. Recently awarded a Marie Curie fellowship, she will work closely with developmental biologists the following 2 years to elucidate the mechanobiology of joint morphogenesis, and then move back to the UPC in Barcelona to complete the CompLimb project in LaCàN.
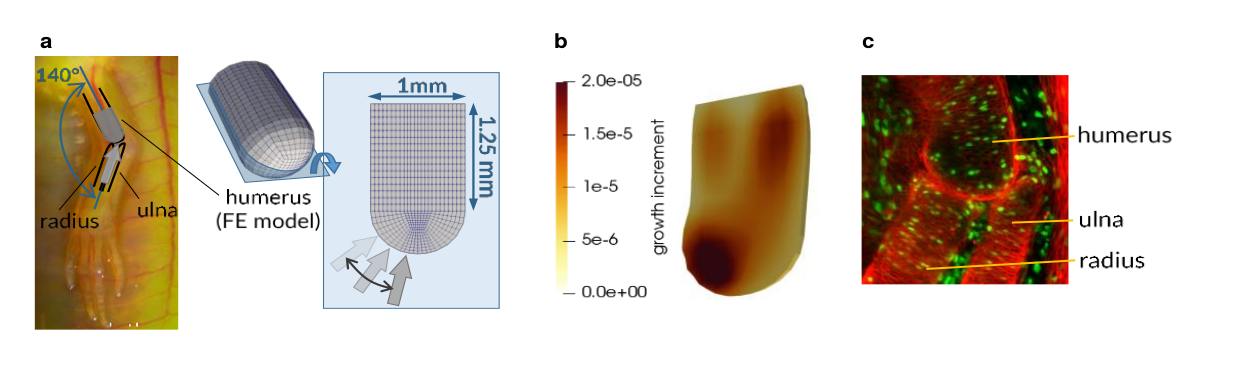
Axolotls regenerate limbs throughout life, providing a powerful animal model for the study of joint development. Axolotl limbs are morphologically similar to human limbs and utilize the same biological rubrics as ontogenic growth. To draw from the therapeutic potential of these similarities, we propose to build a computational model for the prediction of vertebrate limb development. Our model will be based on in vivo data obtained using novel imaging techniques via experiments on axolotl limb growth. In a regenerating axolotl forelimb, we explore the effects of removing mechanical signals (denervation) and transduction of mechanical signals (TRPV4 agonist GSK101). We use click chemistry to co-label and image cell proliferation (alkyne-modified EdU) and matrix (azide-modified methionine) production. Limbs are collected, stained, and imaged with light sheet microscopy, providing a 3D spatial representation of joint structure, cell proliferation, and matrix production. Cell proliferation is reduced in denervated and drug-treated limbs and final joint morphology is altered, supporting the notion that mechanical signals are required for normal limb development. In parallel, we developed a biphasic poroelastic finite element model of growth at tissue level to study how specific changes in limb motion (reduction or removal of mechanical signals) affect joint morphology. A computational model that links the biomechanics and biochemistry of normal and pathological limb development across scales is a powerful predictive tool that can be utilized alongside experiments to determine the physical mechanisms of joint morphogenesis.
